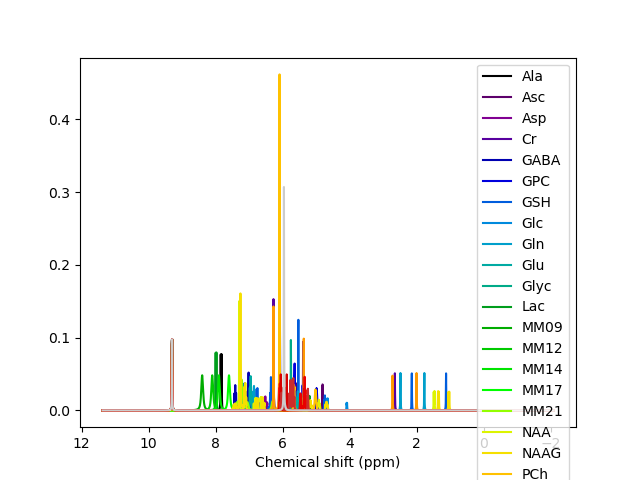
I am trying to use FSL to do a comparison between FSL and Osprey MRS and fMRS processing. We did not collect experimental macromolecule data for our basis set, so I would like to add it into my basis set using the FSL command: “basis_tools add_set --add_MM”. I was successfully able to convert the LCModel .basis to an FSL basis set of .json files.
When I run the command basis_tools add_set --add_MM, the basis set does not visually change, but there are now .json files for each MM in the basis set.
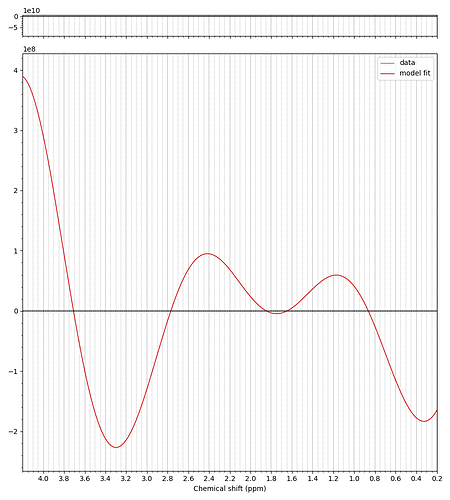
However, when I run the fit with the basis set with added MM, the fit doesn’t work. I assume the fit fails because I am asking it to fit MM that didn’t get correctly added to the basis set.
Regardless, does anyone have suggestions for how to properly add MMs, or why the fit is failing otherwise?
Thank you for any input you can provide!
The following is the fitting call that I have been using:
‘fsl_mrs’,
‘–data’, data_path,
‘–basis’, ‘basis_MM’,
‘–output’, output_path,
‘–h2o’, wref_path,
‘–baseline’, ‘spline, moderate’,
‘–metab_groups’, ‘MM09’, ‘MM12’, ‘MM14’, ‘MM17’, ‘MM21’,
‘–TE’, ‘5’,
‘–TR’, ‘10’,
‘–report’,
‘–overwrite’)
I added the “metab_groups” line only when using the basis with added MM.
I can only add one embed. The fit without MM is a typical fit. The fit WITH MM is nonsense: