I wanted to seek advice on H2O removal during MRS data preprocessing:
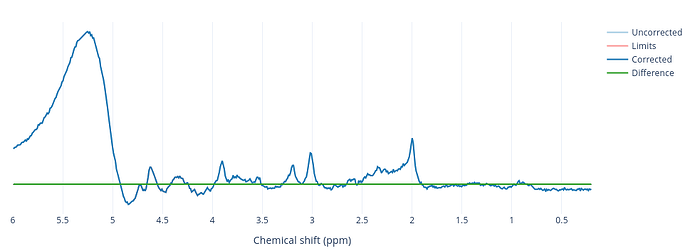
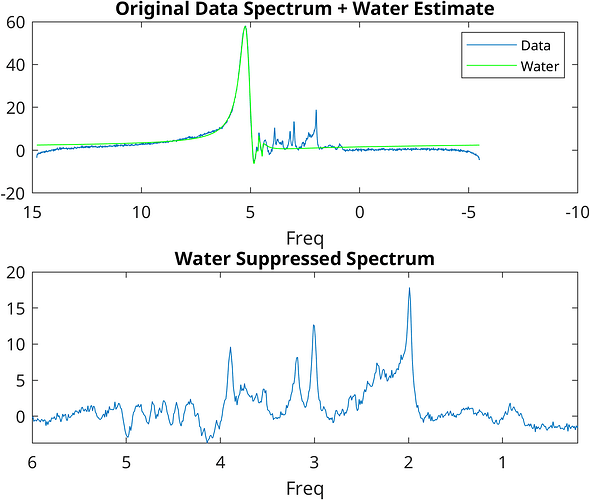
I am using FSL_MRS HLSVD for the residual water removal; however, for certain participants, the water peak is poorly supressed and is Gaussian (possibly movement artifact??), thus I thought of using FID-A for the dominating water peak removal, which has worked well removing the Gaussian peak from 4.3 to 5.2 PPM
Is it okay to use FSL_MRS HLSVD for a few subjects (where only residual water is present) and then FID-A water removal for a few subjects (where water is dominating or Gaussian) in the same experiment?
@ArijitBhattacharya, it’s possible that FSL-MRS is failing to remove the residual water peak using its HLSVD routine because the pre-set frequency range around the water frequency is not broad enough to capture the entire residual peak. But I’ll leave it to @wclarke to confirm.
Hi Both,
Mark you could be right, @ArijitBhattacharya if you click the legend can you see where the limits range is set?
Otherwise I have found some numerical instability in the hlsvdpropy code. It’s soemthing I’ve been meaning to address, but not got around to yet. From memory the roots fed into the vandermonde matrix calculation at hlsvdpropy/hlsvdpropy/hlsvd.py at master · bsoher/hlsvdpropy · GitHub are badly scaled and the whole thing blows up. From memory if you set limits on the roots then this partially fixes it.