Hi Georg,
thanks for your quick response!
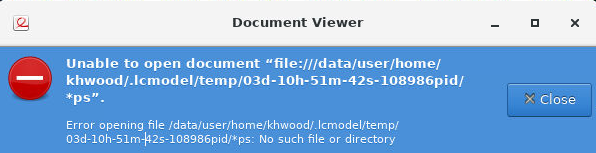
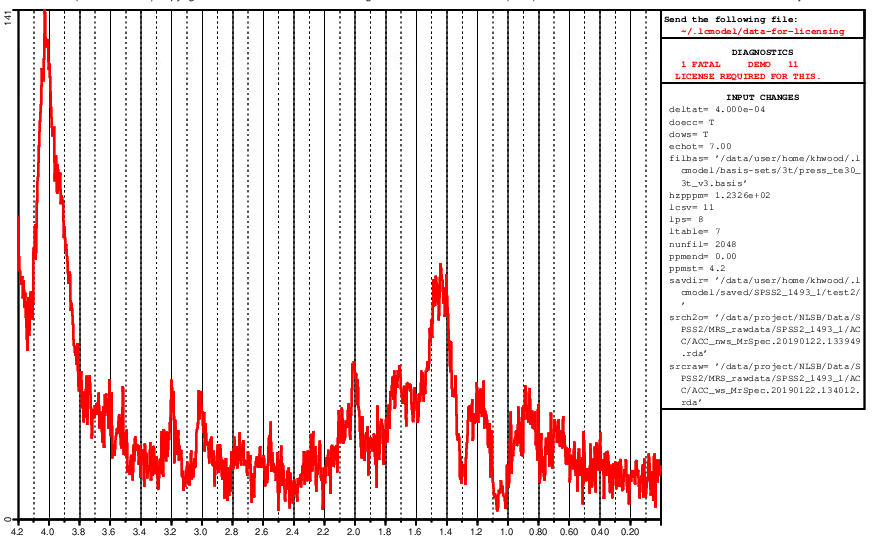
Not exactely, I can select a water reference and after I press ‘ok’ the error is displayed in the bash, it says “LCModel elapsed time = 0 seconds” and evince opens with an error that the .ps cannot be opened which is clear since no file could be created. I hope this makes it clearer where the error happens.
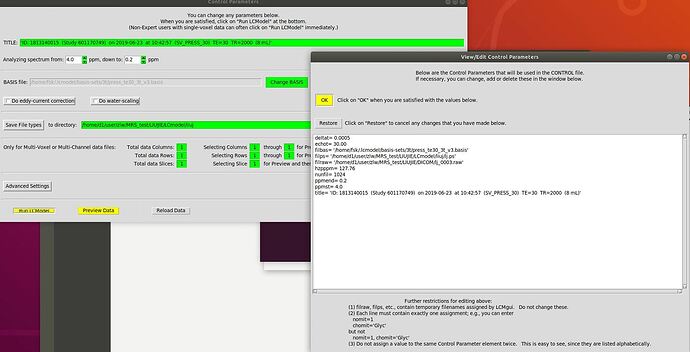
I tried to do so but I am not quite sure which path is needed for the .raw and .ps files since they refer to the temp folder which I guess is created automatically using the GUI. I took the centOS5 control file and exchanged the paths to get this control file:
$LCMODL
title= ‘Soleus’
srcraw= ‘/home/fabian/Dokumente/Files/spectro_MC_04_05_WIP_SV_sLASER_35_10_2_raw_act.SDAT’
srch2o= ‘/home/fabian/Dokumente/Files/spectro_MC_04_05_WIP_SV_sLASER_35_10_2_raw_ref.SDAT’
sptype= ‘muscle-5’
savdir= ‘home/fabian/Dokumente/Files/saved/’
ppmst= 3.8
ppmend= -1.0
nunfil= 1024
ltable= 7
lps= 8
lprint= 6
lcsv= 11
lcoraw= 10
lcoord= 9
hzpppm= 1.2774e+02
filtab= ‘/home/fabian/.lcmodel/temp/06d-21h-53m-00s-0000pid/table’
filraw= ‘/home/fabian/.lcmodel/temp/06d-21h-53m-00s-0000pid/met/RAW’
filps= ‘/home/fabian/.lcmodel/temp/06d-21h-53m-00s-0000pid/ps’
filpri= ‘/home/fabian/.lcmodel/temp/06d-21h-53m-00s-0000pid/print’
filh2o= ‘/home/fabian/.lcmodel/temp/06d-21h-53m-00s-0000pid/h2o/RAW’
filcsv= ‘/home/fabian/.lcmodel/temp/06d-21h-53m-00s-0000pid/spreadsheet.csv’
filcor= ‘/home/fabian/.lcmodel/temp/06d-21h-53m-00s-0000pid/coraw’
filcoo= ‘/home/fabian/.lcmodel/temp/06d-21h-53m-00s-0000pid/coord’
filbas= ‘/home/fabian/.lcmodel/test/output/test.basis’
echot= 42.00
dows= T
deltat= 5.000e-04
lcsi_sav_1 = 12, filcsi_sav_1 = ‘/home/fabian/.lcmodel/temp/06d-21h-50m-00s-0000pid/filcsi_sav_1’, lcsi_sav_2 = 13, filcsi_sav_2 = ‘/home/fabian/.lcmodel/temp/06d-21h-53m-00s-0000pid/filcsi_sav_2’
$END
With this control file I get the following error, which is probably a result of my wrong raw/ps path to the temp folder:
FATAL ERROR ZEROVX 3. (Check LCModel Manual). ************************************************************************
FATAL ERROR MAKEPS 2. (Check LCModel Manual). ************************************************************************
*** FATAL ERROR MAKEPS 2
This error occurred before the plot could be produced.
See the Diagnostics list in the LCModel Manual. ***
*** FATAL ERROR MAKEPS 2
This error occurred before the plot could be produced.
See the Diagnostics list in the LCModel Manual. ***
Adjusting the paths with an already created temp-folder from a try using the GUI, leads to the following error message:
Program received signal SIGSEGV: Segmentation fault - invalid memory reference.
Backtrace for this error:
*** Error in `/home/fabian/.lcmodel/bin/lcmodel’: malloc(): memory corruption: 0x0000000016c42f60 ***
Program received signal SIGABRT: Process abort signal.
Backtrace for this error:
Program received signal SIGABRT: Process abort signal.
Backtrace for this error:
Speicherzugriffsfehler (Speicherabzug geschrieben)
Best
Fabian