Hello!
I’m trying to better understand how the different spectral alignment options affect my GABA peak. For some datasets, the spectra look reasonable when no alignment is applied, but when I use “RestrSpecReg”, the GABA signal develops an unusual shape that persists regardless of the frequency range I specify. This makes me wonder whether I may be applying RestrSpecReg incorrectly, or if there is something specific about these datasets that makes this method problematic. We use the HERMES sequence on 3T, and our files are in TWIX format.
Has anyone encountered a similar distortion of the GABA peak when using RestrSpecReg, or have suggestions for diagnosing what might be causing this?
-
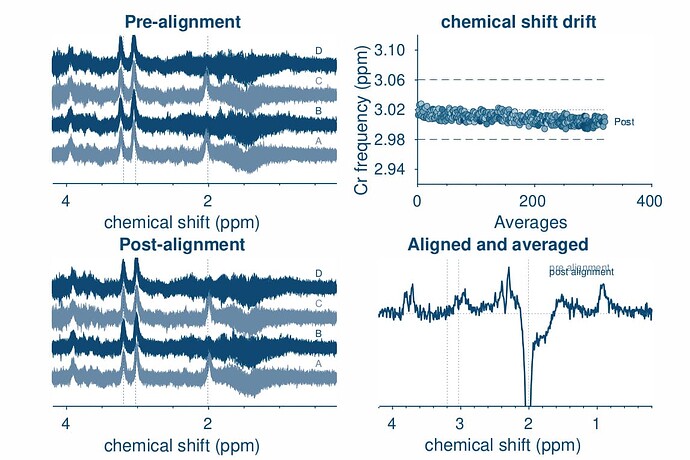
opts.SpecReg = ‘none’;
opts.SubSpecAlignment.mets = ‘L2norm’;
GABA peak looks okay
-
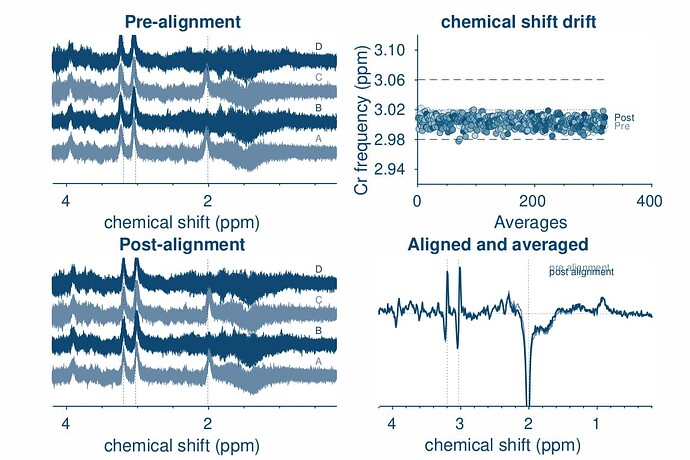
opts.SpecReg = ‘RestrSpecReg’;
opts.SpecRegRange = [1.5 3.5];
opts.SubSpecAlignment.mets = ‘L2norm’;
-
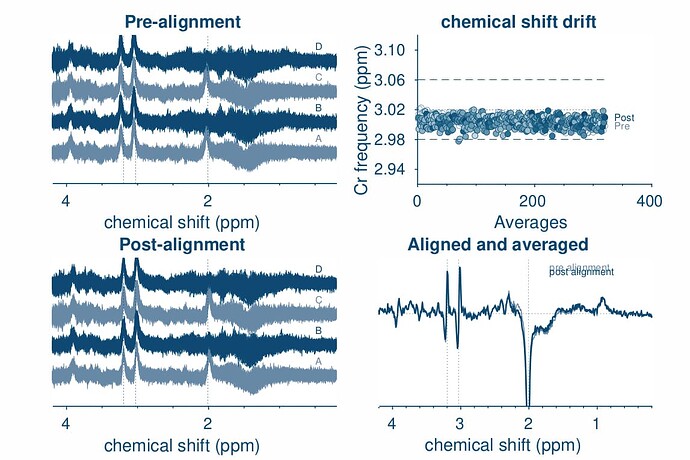
opts.SpecReg = ‘RestrSpecReg’;
opts.SpecRegRange = [2.5 3.5];
opts.SubSpecAlignment.mets = ‘L2norm’;