Hi @alex, I don’t think so since the algorithm processes reconstructed data. However, I have to scrutinize the whole process. @Neda may I know the version of your GE data format?
It’s pfile but I don’t know which what type of X…
Thank you Georg. ![]()
Best,
Dear All,
Hi
I wish you are doing very well.
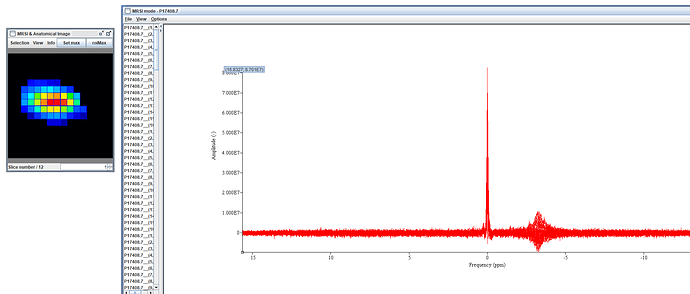
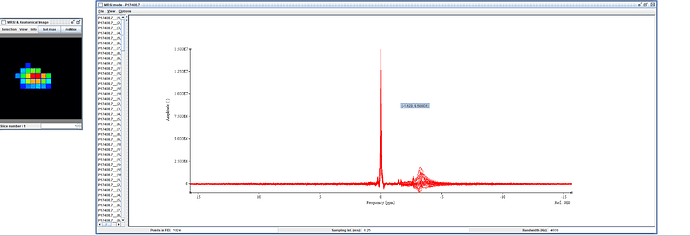
I would like to appreciate Amir for analyzing the data and sending back to me that strange water spectrum there!..I have attached it before and after the coil combination.
There is a question…is it an spurious echo? within the spectrum…
Dear Georg,
Hi,
I wish You could make sometime to pay a look at TARQUIN screenshots I uploaded here…Thank you very much in advance…
They looked fine to me, but I’m not a TARQUIN expert and I haven’t used it in maybe 5 years. @martin is the developer and should be able to advise in greater detail.
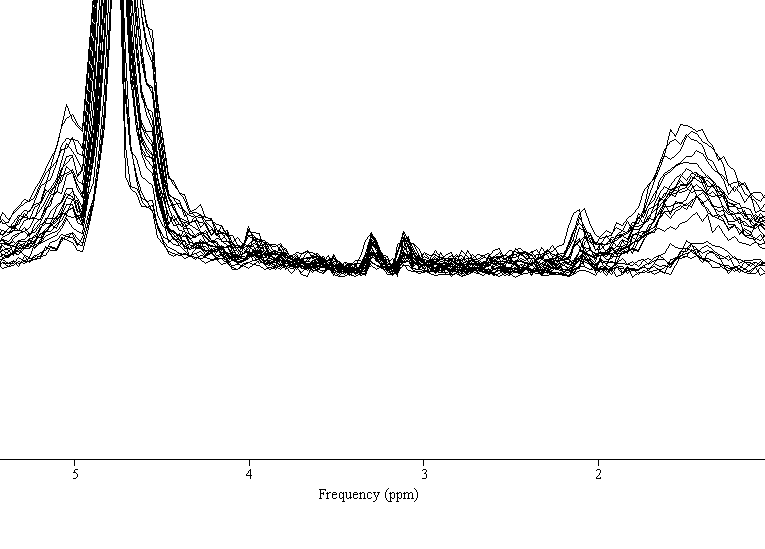
What you see in your jMRUI output are large and highly variable lipid contaminations, not spurious echos. (If you set the frequency axis to have water at 4.68 ppm, they will be between 0 and 2 ppm).
Thank you for following up. Here is a closer look. (Coil-combined and Magnitude)
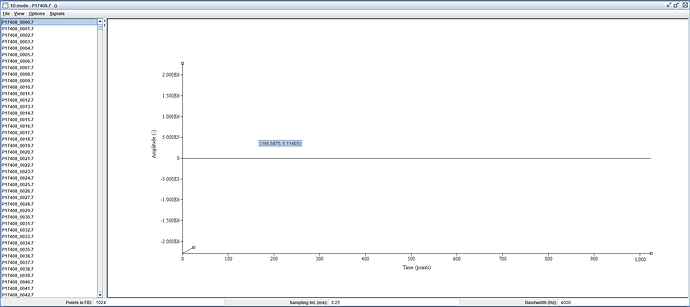
From the header, I can see that a reference scan per coil was acquired (12 spectra). However, these spectra are zero at all time points.
@NNeda Here, They proposed an approach for removing lipid contamination. It might help you.
Thank you very much for your kind attention. ![]()
No. Zero-data will be caused by an acquisition that doesn’t work in the intended way, and it is quite likely that the data acquisition protocol (exam card) needs revision.
What’s your situation on site? Do you have someone who is responsible for MRS data acquisition?
Yes, There is MR technologist . I analyzed a single voxel data from our scanner and it reported the water signalcorrectly…Its very likely that it’s not acquired or saved in CSI acquisition mode o …but I’m not certain about it yet as it displays the FWHM and suppression efficiency while prescanning. Do you mean adding it to the protocol?..It’s only possible if purchased for the scanner…right?..
I would highly recommend that you contact someone who has successfully worked with your MRSI sequence before. It is not trivial to set up an MRSI protocol correctly, and I don’t think we can solve it from a distance without access to the full protocol, hardware setup, software versions, etc. I think @amirshamaei and @alex may have experience with it. @noeskera might have a decent idea to point you towards someone, since he has the distinguished title of “Global Spectroscopy Leader” at GE.
Yes you are right. .Thanks alot for your advice…As Alex said before water signal was not a necessity for my case but it turned out to be an interesting question about what really happens to it during GE MRSI acqusition…
Hi, this is Ralph, the “GE Global Spectroscopy Leader” as Georg mentioned ![]() I am “just” leading the MRS developments in GEHC, so enough space for others to really “lead” MRS in general. But now to the issue. I assume you are using the product CSI sequence (PROBE-P). Other than in SV where we acquire non-water suppressed reference data automatically and store this data in the beginning of the pfile, we are not acquiring reference data in CSI. This would double the acquisition time and therefore you need to do this manually (unfortunately you need to be in research mode to do this as you manually need to switch off water suppression by a CV “suppress”). To my knowledge GE pfile MRSI support in jMRUI wasn’t really working until recently Ana Jorge Goncalves added this during her PhD thesis. Not sure if that is already part of the official jMRUI release. Let me know if you need more information. Cheers
I am “just” leading the MRS developments in GEHC, so enough space for others to really “lead” MRS in general. But now to the issue. I assume you are using the product CSI sequence (PROBE-P). Other than in SV where we acquire non-water suppressed reference data automatically and store this data in the beginning of the pfile, we are not acquiring reference data in CSI. This would double the acquisition time and therefore you need to do this manually (unfortunately you need to be in research mode to do this as you manually need to switch off water suppression by a CV “suppress”). To my knowledge GE pfile MRSI support in jMRUI wasn’t really working until recently Ana Jorge Goncalves added this during her PhD thesis. Not sure if that is already part of the official jMRUI release. Let me know if you need more information. Cheers
Dear Ralph,
Hi
Thank you very much for your complete explanation.Now, everything is clear… ![]()
![]()
Hi NNeda
Can I analyze 3D (Multivoxel) MRSI data with Tarquin software?
Hi dear researchers
I have a technical question please guide me:
Can I analyze 3D (Multivoxel) MRSI data from GE MRI 360 (Pfile) with Tarquin software?
Hi Javid
I tested a 3D MRSI pfile from GE MR Discovery 750 by TARQUIN 11 and 10 but unfortunately it stopped while preprocessing…it seems there is a problem with 3D pfile on TARQUIN…
Dear Ralph,
Hi
Is the water FWHM displayed in GE spectroscopy while prescanning, representative of global shimming?
Thank You Very Much