Hi!
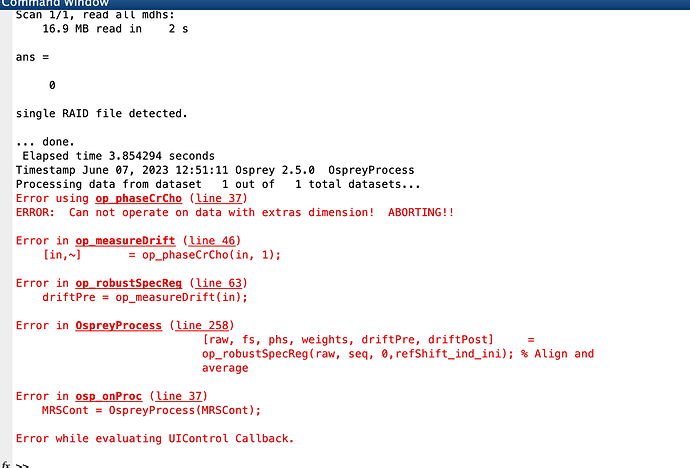
I’m trying to re-analyze a published dataset (Kurcyus et al 2018 https://doi.org/10.1523%2FJNEUROSCI.1214-18.2018)using Osprey and I’m running into trouble while trying to process the data. Both using the entire dataset as well as for single subjects, the data loads fine, but at the processing stage it seems to find too many dimensions in the op_phaseCrCho.m function. I have attached the job file for reference.
jobTwix_RP_try.m (17.1 KB)
Thanks in advance!
Hi @RashiPant,
This error gets thrown if: in.dims.extras>0. Which MEGA-PRESS sequence is this? Perhaps the subspectra are getting, or an additional singleton dimension is not being collapsed during Osprey’s load routines.
Could you please check the dims structure, and also the shape of your fids/specs arrays?
If you can share the data, I’m also happy to take a look.
Cheers,
Chris
Hi Chris, thank you so much for getting back to me!
The data was collected using a Siemens twix MEGA-PRESS sequence (WIP #795 package) with a Cr reference. I can’t seem to get any output in the MRS_container from Osprey - The MRS_Struct structure for Gannet, which seemed to run alright, has 2 subspectra and the following fids structure:
MRS_Struct.fids
ans =
struct with fields:
data: [2080×256 double]
ON_OFF: [1×256 double]
data_align: [2080×256 double]
I’m attaching a link to the data as well - really sorry for the trouble!!
https://drive.google.com/drive/folders/11Q1Mq6t4irPaSQAwGa9mT_OhhZRsO0Fo?usp=drive_link
Hi @RashiPant,
I had a look at the data. It seems that Osprey is misattributing the subspec dimensions for this particular version of the WIP sequence. I’ve created an issue on Github. I have a workaround running on my local machine, and I’ll try to push a fix before the weekend.
Best,
Chris
Hi @RashiPant ,
I pushed a small fix to the develop branch of Osprey on Github. This works for the example data you sent me. Try pulling the latest version and let me know if you run into any issues.
Cheers,
Chris
Thank you so, so much!! Trying it out now
Worked perfectly aahh!!! Thank you!
Hi @Chris_Davies-Jenkins,
I have got the same issue but with the latest version of Osprey. The MEGA-PRESS TWIX data were acquired using XA50. Would you mind to take a look of my data?
Thanks,
Steve
Problem solved by adding XA50 version in io_loadspec_twix.m
1 Like