Ok, The reason i highlighted Ivan’s work: I only tailored them to work with B1 shimming and/or Nova coil. Once you setup these things, it works in almost every brain region without having any problem. So i need to highlight this one more time, the recipe is on the following paper.
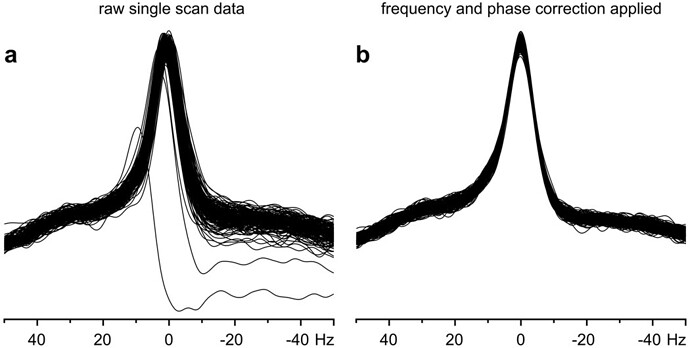
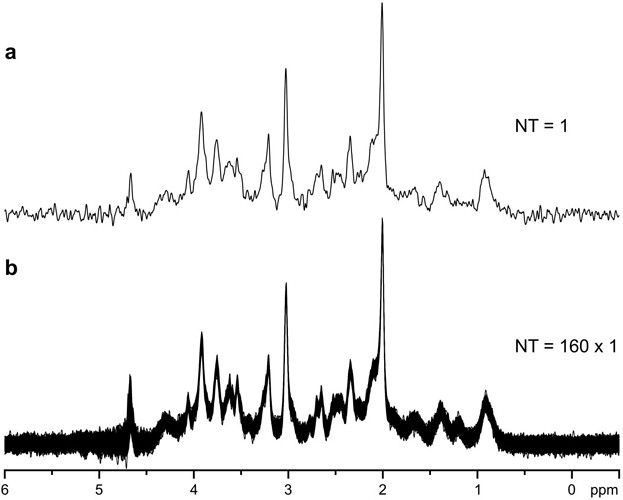
“Good localization performance and excellent water suppression resulted in highly reproducible single-scan spectra, without any significant contamination by unwanted coherences. This makes the sequence much more robust in comparison to sequences that require phase cycling to subtract unwanted coherences. When depending on phase cycling for the elimination of such signals, instabilities during an experiment, e.g., physiological motion of a subject, can result in subtraction errors and insufficient suppression of unwanted signals.To achieve the highest possible spectral resolution, elimination of all effects that unnecessarily increased the line width was desirable. Physiological motion, such as breathing, can cause periodic frequency variations up to ±2 Hz at 7 T. In addition, small random motions of the head during the study can induce phase fluctuations. Both effects were easily eliminated when single-scan data averaging was used (Fig. 2).”
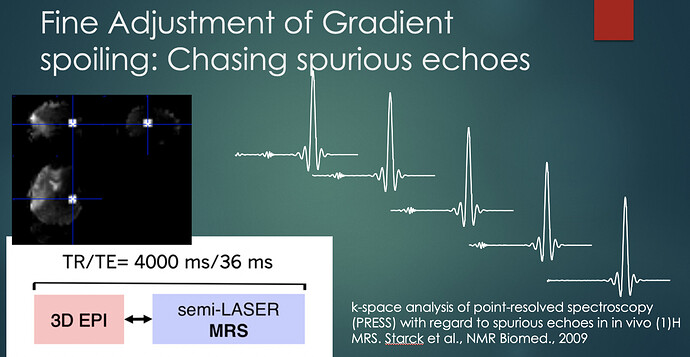
There are two problems with OVS. One is it increases the SAR. But if you are using VAPOR then you do not need to worry about TR so it is not a problem anymore. The other one is the distance between OVS slabs and VOI. Ivan set this as 7mm and I tested them anything less than 7mm will result in signal suppression. I did have the illustration regarding this but I could not find it. Now if your lipid signal falls in between this region, then your VOI profile will bring it back to your spectra. This is a problem when you are working on occipital cortex ROI for fMRS. DOTCOPS or the following paper might solve the problem. But to be honest there is not any problem once you do shim and place the voxel very well.
https://onlinelibrary.wiley.com/doi/full/10.1002/mrm.26213
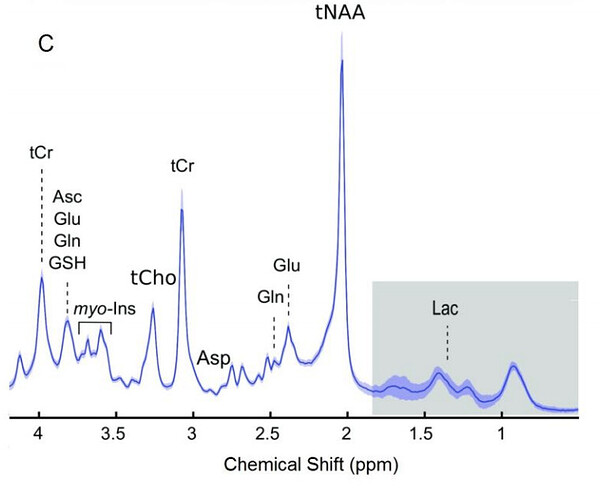
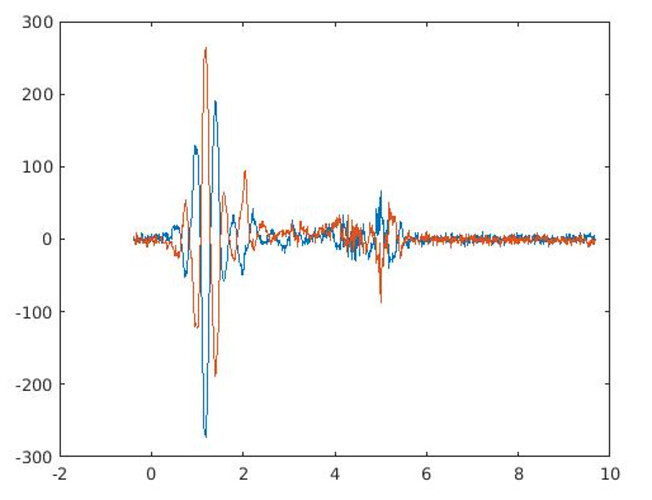
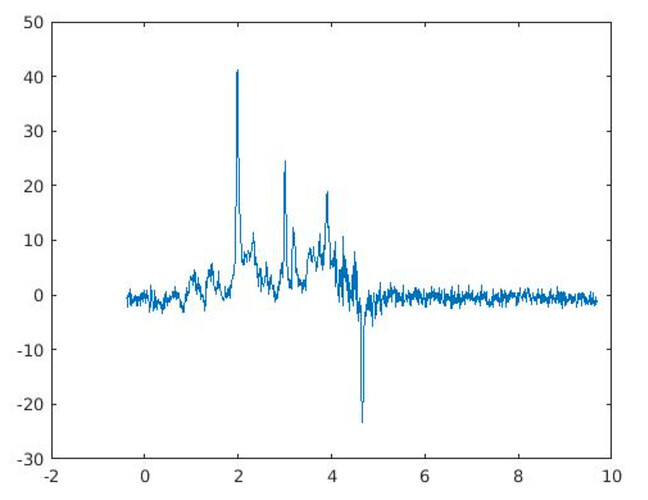
In my first OxfordWIN/fMRIB fMRS paper, we did have the problem of negative/positive peaks between 0.9 and 2 ppm, which was not terrible. Since I did not have the luxury of using CMRR hardware options, I was limited to standard hardware. Thus in the following paper, we used a limited ppm range.
https://www.cell.com/current-biology/pdf/S0960-9822(15)00434-0.pdf
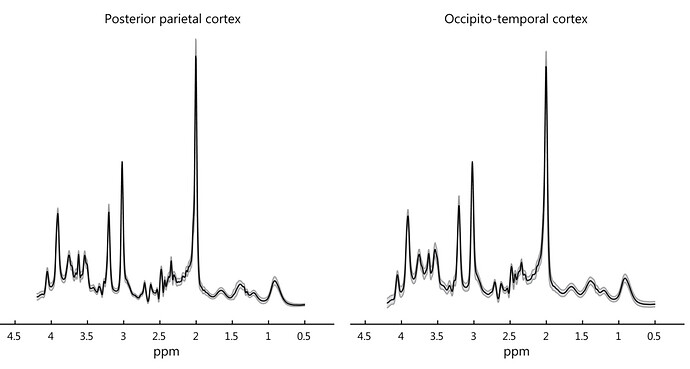
But I was not happy about this and I spent quite some time on RF pulse orders, gradients schemes since there was and is still a lot of demand at OxfordWIN/fMRIB for fMRS works. Then I reshuffled everything. CMRR spin-echo adiabatic (SEAD, , semi-LASER) sequence was using excitation in the X direction and their VAPOR-OVS partners (crusher gradients and pulses together). I realized with the NOVA coil this problem is originating from this order. then i change the orientation from Transverse to Coronal which resulted in a very reproducible spectral pattern in almost every brain region.
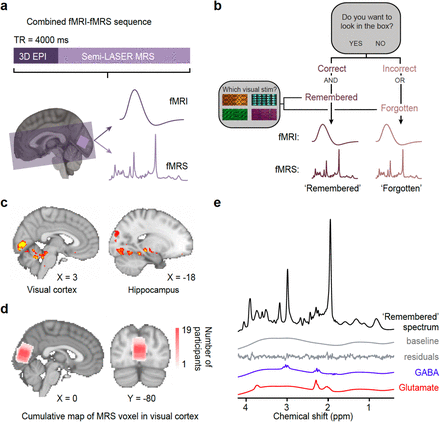
https://www.nature.com/articles/s41467-019-08313-y#MOESM1
Thus the problem you highlighted was not there anymore maybe a little bit in the ACC voxel.
I have done these in 2013/4 and I have not run any fMRS study and analyze them by myself. Thus Another trick to having a successful fMRS study is having excellent collaborators(Betina, Hellen, Zoe, Charlie, Holly… ) who understand/value MRS very well. Also, radiographers should also be on this team.
For instance, I have not acquired, analyzed, and reported any of these fMRS studies, I only looked at the results at the final stage. This way I minimized the possible bias that might originate from me. So most of them are blind design studies for me and we identify the analysis parameters in advance. Also, I have not acquired those data, this also minimizes any biases. For this, I have to thank OxfordWIN/FMRIB radiographers since they trusted me and followed the steps I suggested.
Overall, I think special thanks go to Ivan’s works.

 .
.