Dear Dr. Oeltzschner and al.,
I am fairly new to MRS, but your software has been of great help for me lately as I started processing data for one of my graduate projects. So far I have worked with Siemens TWIX, DICOM and RDA file formats. Everything has worked very nicely (especially for TWIX data) and I look forward to continue using your software. Below is a list of questions I have, and possible bugs that I have encountered - I’m not sure if this is the best forum for this, so if I need to post somewhere else please do let me know.
-
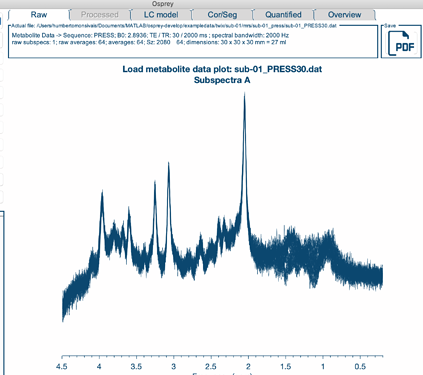
I noticed that after the latest update, the ‘Processed’ tab in the GUI is grayed out when loading a MRSCont.mat file of data that has already been processed. This happens after I have gone through all of the processing steps and I want to review the data again or save different pdfs. I can navigate through all the other tabs except for the ‘Processed’ tab (see screenshot below).
-
When saving pdfs of the ‘Cor/Seg’ tab, the voxel and voxel fraction information does not show up in the pdf. Will this information show if this specific module is plotted independently? Does it need to be saved as a png or jpeg instead?
-
I used the generated RDA files for fitting and quantification with LCModel and noticed that a lot of the header information is missing, i.e., patient id, sequence description, etc… This doesn’t interfere with the fitting or quantification, but some of title information has to be entered manually.
-
I tried using the LCModel control files in the output folder and kept getting errors. Is there something I need to change within the control file? Like the paths?
-
Lastly, is there a feature to import existing LCModel basis sets into Osprey format? I would like to compare some of the fitting and quantification results using Osprey and LCModel for the sLASER sequence.
Thank you and I appreciate any answer you could provide.
-Humberto M.